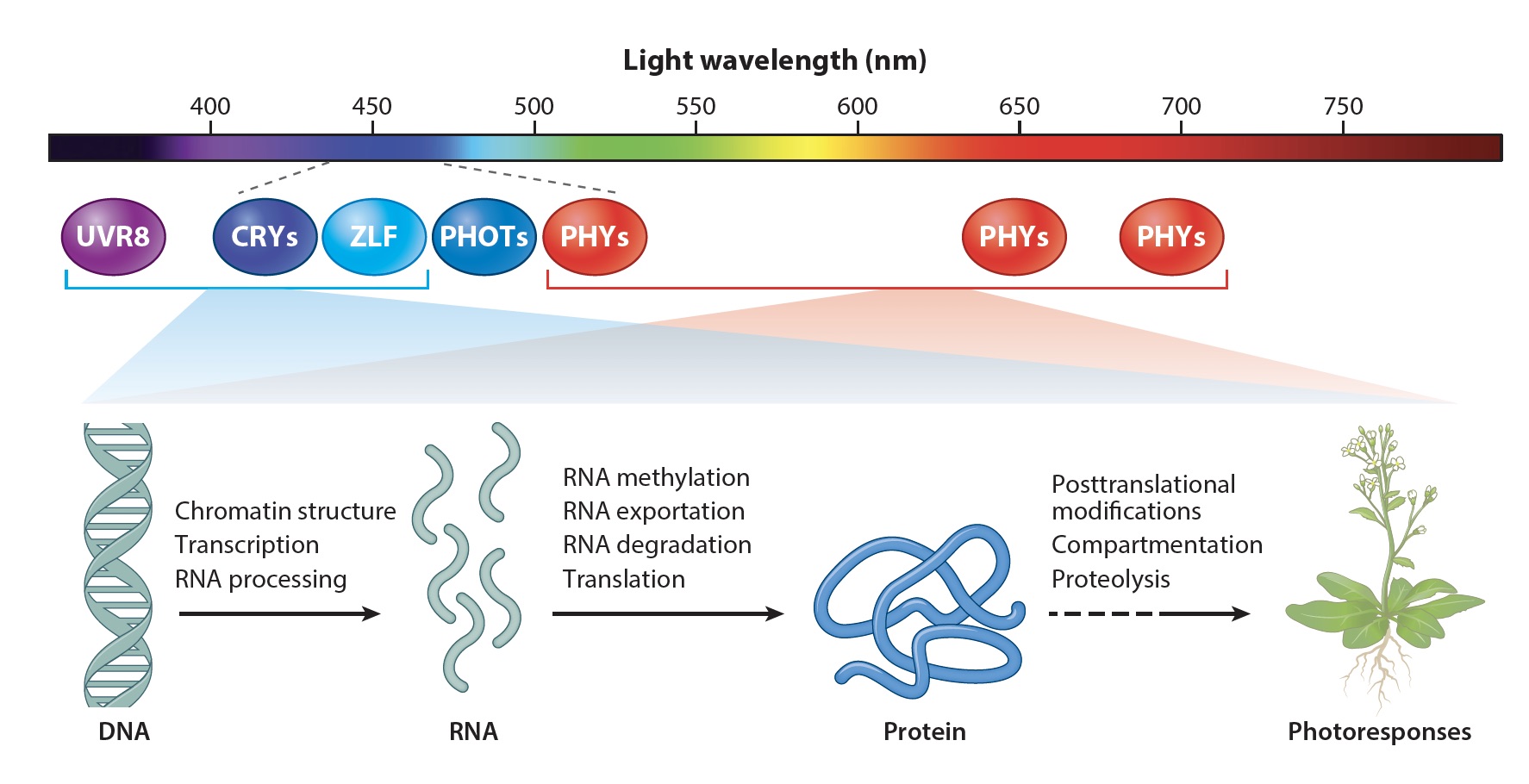
Fig. 1. Plant Photorecptors
Research Interests
My lab uses molecular genetics, biochemistry, and genomics approaches to study plant photoresponses and the molecular mechanisms of cryptochromes in the model plant Arabidopsis. Cryptochromes are ancient photoreceptors that widely found in microbials, plants, and animals, including human. Cryptochromes act as blue-light receptors to mediate photoresponses of plants. The genomes of most land plants encode two clades of cryptochromes, CRY1 and CRY2, which are nucleocytoplasmic proteins that regulate genomic expression in response to blue light. Photoresponsive protein-protein interaction is the primary mode of signal transduction of cryptochromes. Cryptochromes exist as physiologically inactive monomers in the dark; the absorption of photons leads to conformational change and cryptochrome homooligomerization, which alters the affinity of cryptochromes interacting with cryptochrome-interacting proteins to form various cryptochrome complexes. These cryptochrome complexes, collectively referred to as the cryptochrome complexome, alter transcription, RNA methylation, RNA splicing, protein modification, and protein degradation to modulate plant growth and development. The activity of cryptochromes is regulated by photooligomerization; dark monomerization; cryptochrome regulatory proteins; and cryptochrome phosphorylation, ubiquitination, and degradation. Most of the more than 30 presently known cryptochrome-interacting proteins are either regulated by other photoreceptors or physically interacting with the protein complexes of other photoreceptors. Some cryptochrome-interacting proteins are also hormonal signaling or regulatory proteins. These two mechanisms enable cryptochromes to integrate blue-light signals with other internal and external signals to optimize plant growth and development.
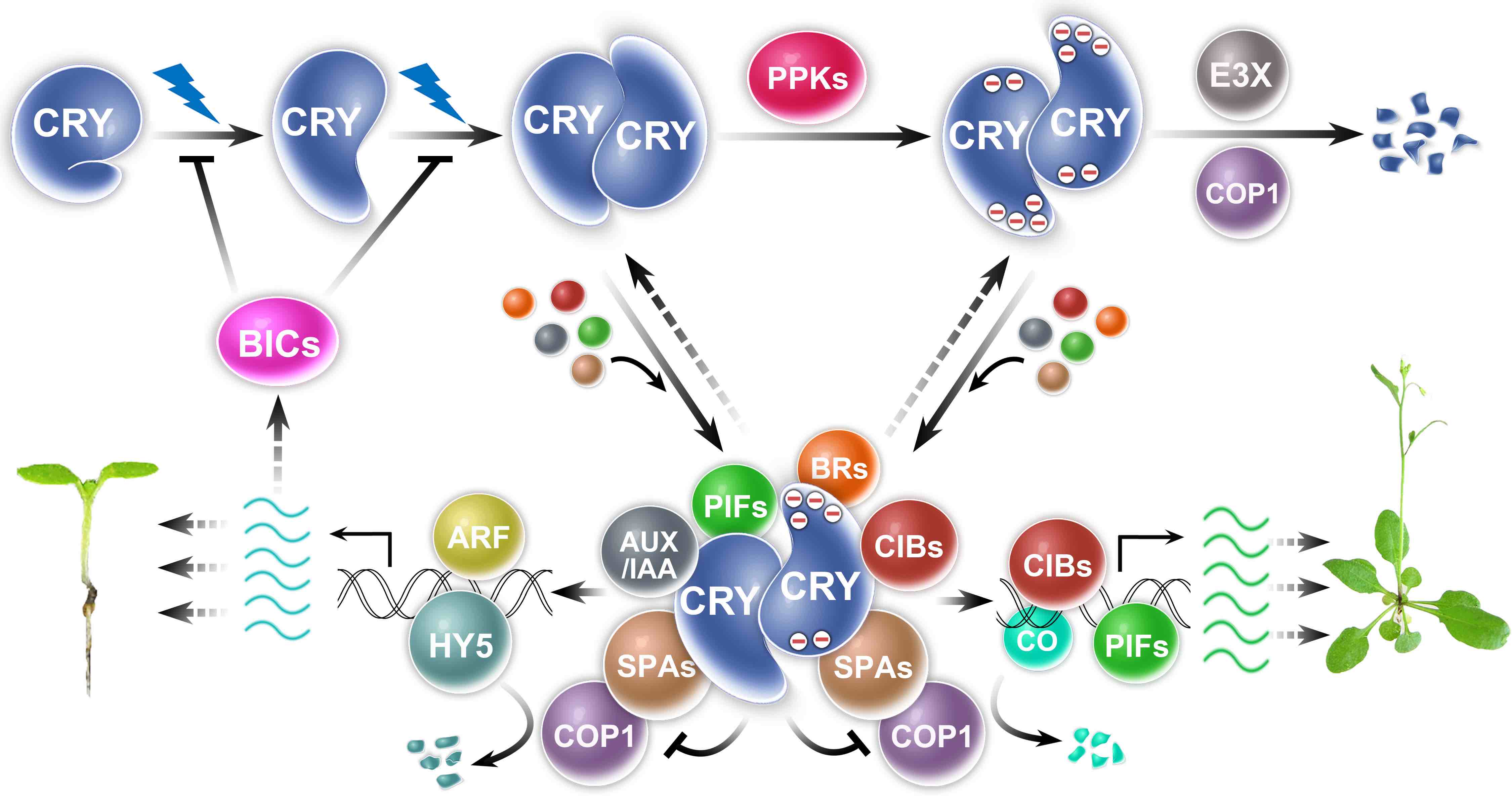
Fig. 2. How cryptochromes work
References:
- Wang, Q., and Lin, C. (2020) Mechanisms of cryptochrome-mediated photoresponses in plants. Annual Review of Plant Biology 71:23.1-23.27.
- Liu, Q., Wang, Q., Deng, W., Wang, X., Piao, M., Cai, D., Li, Y., Barshop, W. D., Yu, X., Liu, B., Oka, Y., Wohlschlegel, J., Zuo, Z., and Lin, C., (2017), Molecular basis for blue light-dependent phosphorylation of Arabidopsis cryptochrome 2, Nature Communications 8: 15234.
- Wang, Q., Zuo, Z., Wang, X., Gu, L., Yoshizumi, T., Yang, Z., Yang, L., Liu, Q., Liu, W., Han, Y.-J., Kim, J.-I., Liu, L., Wohlschlegel, J.M., Matsui, M., Oka, Y., and Lin, C. (2016) Photoactivation and inactivation of Arabidopsis cryptochrome 2. Science 354:343-347.
- Liu, H. Yu, X., Li, K., Klejnot, J., Yang, H., Lisiero, D. and Lin, C. (2008) Photoexcited CRY2 interacts with CIB1 to regulate transcription and floral initiation in Arabidopsis. Science 322: 1535-1539
- Lin, C. and Shalitin, D. (2003) Cryptochrome structure and signal transduction. Annu. Rev. Plant Biol. 54: 469-496.
- Shalitin, D., Yang, H., Mockler, T. C., Maymon, M., Guo, H., Whitelam, G. C., and Lin, C. (2002) Regulation of Arabidopsis cryptochrome 2 by blue light-dependent phosphorylation. Nature 417:763-767
- Guo, H., Yang, H., Mockler,T. C., and Lin, C. (1998) Regulation of Flowering Time by Arabidopsis Photoreceptors. Science 279:1360-1363.